10 Unsupervised Machine Learning Algorithms: What Are They And How To Create Them
Machine Learning is one of the hottest software development topics right now. The algorithms and techniques which enable machine learning have begun to really mature and have graduated from ‘interesting ideas’ into providing genuine power and permitting capabilities in our apps which can sometimes seem magical just as much as they are useful. Python has very quickly emerged as a de facto language for machine learning. There is rich set of machine learning libraries available for Python providing the ability to do everything from image recognition to complicated scientific analyses.
How to use machine learning libraries to Delphi
Did you know that it’s simple to use some truly excellent Python libraries to boost your Delphi app development on Windows? All that wonderful array of Python machine learning goodness is also easily available to you as a Delphi programmer.
Adding Python to your Delphi code toolbox can enhance your app development by bringing in new capabilities that allow you to provide innovative and powerful solutions to your app’s users, combining the best of Python with the supreme low-code and unparalleled power of native Windows development that Delphi provides.
Are you looking for how to build a GUI for a powerful Unsupervised Machine Learning library?
You can build a state-of-the-art unsupervised learning solution with scikit-learn on Delphi. This article will demonstrate how to create a Delphi GUI app dedicated to the scikit-learn library.
Watch this video by Jim McKeeth for a thorough explanation of why you can love both Delphi and Python at the same time:
What is the scikit-learn machine learning library?

scikit-learn is an open-source Python machine learning library. scikit-learn has simple and efficient tools for predictive data analysis that are built on top of SciPy, NumPy, and Matplotlib.
Support vector machines, random forests, gradient boosting, k-means, and DBSCAN are among the algorithms available in scikit-learn for classification, regression, and clustering.
In this article, we will specifically talk about clustering algorithms.
What is unsupervised machine learning?
Unsupervised learning, also known as unsupervised machine learning, analyzes and clusters unlabeled datasets using machine learning algorithms.
Without human intervention, these algorithms uncover hidden patterns or data groupings. Its ability to detect similarities and differences in data makes it an ideal solution for exploratory data analysis, cross-selling strategies, customer segmentation, and image recognition (source: IBM Cloud Education, 2020).
What is clustering and how does it relate to machine learning?
Clustering is a type of unsupervised learning problem. Cluster analysis is another name for this technique.
It is frequently used as a data analysis technique for discovering interesting patterns in data, such as customer groups based on their behavior.
There are numerous clustering algorithms available, and there is no single best clustering algorithm for all cases. Instead, it is a good idea to experiment with various clustering algorithms and different configurations for each algorithm.
A cluster is frequently a dense area in the feature space where examples from the domain (observations or rows of data) are closer to the cluster than to other clusters. The cluster may have a sample or point feature space as its center (the centroid), as well as a boundary or extent (source: Brownlee, machinelearningmastery.com, 2020).

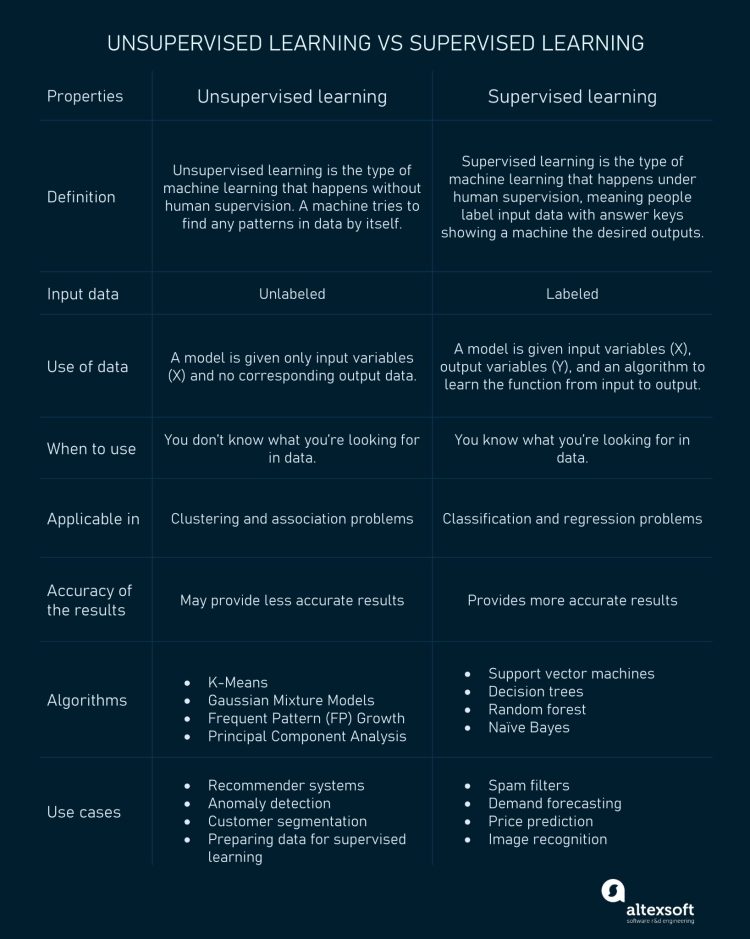
What is the difference between supervised and unsupervised machine learning?
The following infographic created by AltexSoft may give you quick insights into the differences between supervised and unsupervised learning:
How do I get and install the scikit-learn library?
You can easily install scikit-learn with pip:
|
pip install –U scikit–learn |
Or, if you are using Anaconda Python distribution, you can use this command to avoid complexities and conflicts between required libraries:
|
conda install –c anaconda scikit–learn |
And usually, scikit-learn is already installed by default if you use the Anaconda Python distribution.
How do I install Python4Delphi?
Follow the Python4Delphi installation instructions mentioned here. Alternatively, you can follow the simple instructions in Jim McKeeth‘s “Getting Started With Python4Delphi” video:
How do I build a Delphi GUI for the scikit-learn machine learning library?
Our project’s user interface structure is as follows:

Here is the list of Components used in the fastai4D demo app:
- TPythonEngine
- TPythonModule
- TPythonType
- TPythonVersions
- TPythonGUIInputOutput
- TForm
- TMemo
- TOpenDialog
- TSaveDialog
- TSplitter
- TImage
- TPanel
- TLabel
- TComboBox
- TButton
Navigate to the UnitScikitLearn4D.pas, and add the following line to the FormCreate, to load our basic scikitlearnApp.py:
|
Memo1.Lines.LoadFromFile(ExtractFilePath(ParamStr(0)) + ‘scikitlearnApp.py’); |

And make sure that the scikitlearnApp.py is in the same directory as our ScikitLearn4D.exe or inside your Delphi project folder.
You can change the “scikitlearnApp.py” with any scikit-learn script you want, or you can load your scikit-learn scripts at runtime, by clicking the “Load script…” like we will show you in the next Demo Sections.
How do I use scikit-learn to run machine learning clustering algorithms in a Delphi app?
This post will walk you through the process of integrating Python scikit-learn into a Delphi-created Windows app. As an added bonus, we will compare ten different clustering algorithms.
Highly recommended practice:
1. This GUI was created by modifying Python4Delphi Demo34, which allows us to change the Python version in the runtime (this will save you from the seemingly complicated dll issues).
2. Add “Jpeg” to the top of our UnitScikitLearn4D.pas code’s Uses-list. We must do so because Delphi will not understand the JPG format otherwise. It should work with this change.
After that, the above modification should look like this:

We can also load JPG images into our TImage.
3. Set up the following paths in your Environment Variable:
|
C:/Users/ASUS/AppData/Local/Programs/Python/Python38 C:/Users/ASUS/AppData/Local/Programs/Python/Python38/DLLs C:/Users/ASUS/AppData/Local/Programs/Python/Python38/Lib/bin C:/Users/ASUS/AppData/Local/Programs/Python/Python38/Lib/site–packages/bin C:/Users/ASUS/AppData/Local/Programs/Python/Python38/Scripts |
Set up the following paths if you’re using the Anaconda Python distribution:
|
C:/Users/ASUS/anaconda3/DLLs C:/Users/ASUS/anaconda3/Lib/site–packages C:/Users/ASUS/anaconda3/Library C:/Users/ASUS/anaconda3/Library/bin C:/Users/ASUS/anaconda3/Library/mingw–w64/bin C:/Users/ASUS/anaconda3/pkgs C:/Users/ASUS/anaconda3/Scripts |
4. To show plotting results, you must use Matplotlib outside of the “normal” command-line process. To do this, add the following lines to all of your Python code:
|
import matplotlib import matplotlib.pyplot as plt matplotlib.use(“Agg”)
…
plt.savefig(“scikitlearnImage.jpg”) |
We strongly advise you to name your image output as “scikitlearnImage.jpg” in order for it to load automatically on your GUI after you click the “Show plot” button.
5. Set MaskFPUExceptions(True); to the UnitScikitLearn4D.pas file to avoid Delphi raising an exception when floating operations result in +/- infinity (e.g. division by zero), which is caused by incompatibility with a number of Python libraries such as NumPy, SciPy, pandas, and Matplotlib.
One of the best features of this ScikitLearn4D Demo GUI is that you can choose your preferred Python version, which is interchangeable.
We tested this ScikitLearn4D GUI for both regular Python and Anaconda Python, and it works best for both Python versions.
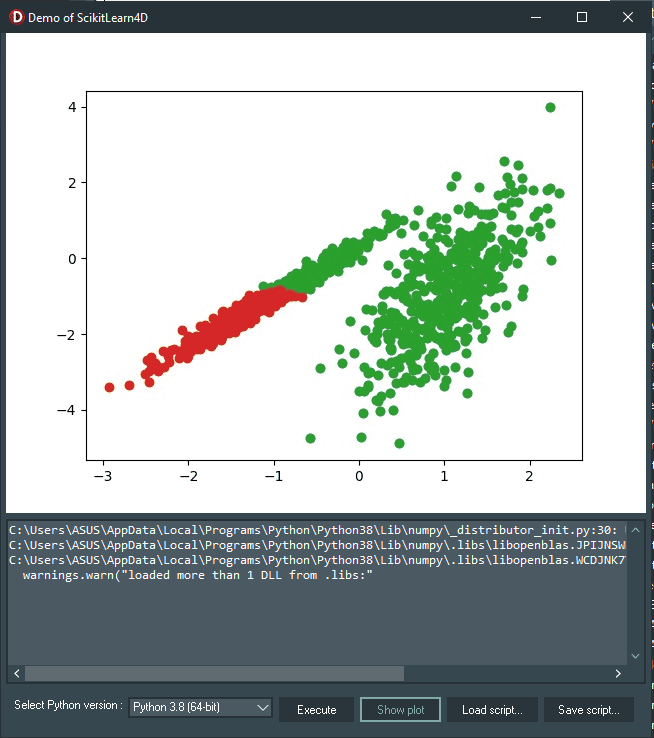
Then, click the “Execute” button to run the very basic example to create and plot a test binary classification dataset for this clustering demo (the Python code is already called inside the UnitScikitLearn4D.pas file), and then click the “Show plot” button to display the figure. Here is the result:

How to create a clustering dataset using scikit-learn on the Delphi app?
In this article, we will use the make_classification() function to create a test binary classification dataset.
There will be 1,000 examples in this dataset, with two input features and one cluster for each class. Because the clusters are visually obvious in two dimensions, we can plot the data with a scatter plot and color the points in the plot according to the assigned cluster. This will enable us to see how “well” the clusters were identified, at least on the test problem.
This test problem’s clusters are based on a multivariate Gaussian, and not all clustering algorithms will be effective at identifying these types of clusters. As such, the results in this article should not be used as the basis for comparing the algorithms generally.
To create this dataset, click the “Execute” button and click the “Show plot” button to show the plot as we’ve mentioned in the previous section.

How to perform clustering algorithms using scikit-learn on the Delphi app?
What is affinity propagation?
Quoting Mézard, 2007:
“Affinity propagation is known in computer science as a message-passing algorithm and suggests that the algorithm can be understood by adopting an anthropomorphic viewpoint.
Imagine that each item being clustered sends messages to all other items informing its targets of each target’s relative attractiveness to the sender. Given the attractiveness messages received from all other senders, each target then responds to all senders with a reply informing each sender of its availability to associate with the sender. Senders process the data and respond to the targets with messages informing each target of the target’s revised relative attractiveness to the sender, based on the availability messages received from all targets. The message-passing procedure proceeds until a consensus is reached on the best associate for each item, considering relative attractiveness and availability. The best associate for each item is that item’s exemplar, and all items sharing the same exemplar are in the same cluster. Essentially, the algorithm simulates conversation in a group of people, where each seeks to identify his or her best representative for some function by conversing with all others.”
To try out this Affinity Propagation algorithm example, load the demo01_affinityPropagation.py at runtime by clicking the “Load script…” button, and then “Execute”. Here is the result:

The AffinityPropagation class is used to write the demo01_affinityPropagation.py, and the main configuration to tune is the “damping” set between 0.5 and 1, and perhaps “preference”.
Finally, Affinity Propagation is an unsupervised machine learning algorithm that is particularly well suited for problems in which the optimal number of clusters is unknown. As a result, it performed poorly in this demo.
What is agglomerative clustering?
Quoting Ackermann et. al, 2014:
“Agglomerative clustering is a bottom-up clustering process. At first, each input object forms its own cluster. The two ‘closest’ clusters will be merged in each subsequent step until only one cluster remains.
This clustering process generates a hierarchy of clusters, such that for any two different clusters A and B from possibly different levels of the hierarchy, we have either A ∩ B = ∅, A ⊂ B, or B ⊂ A.“
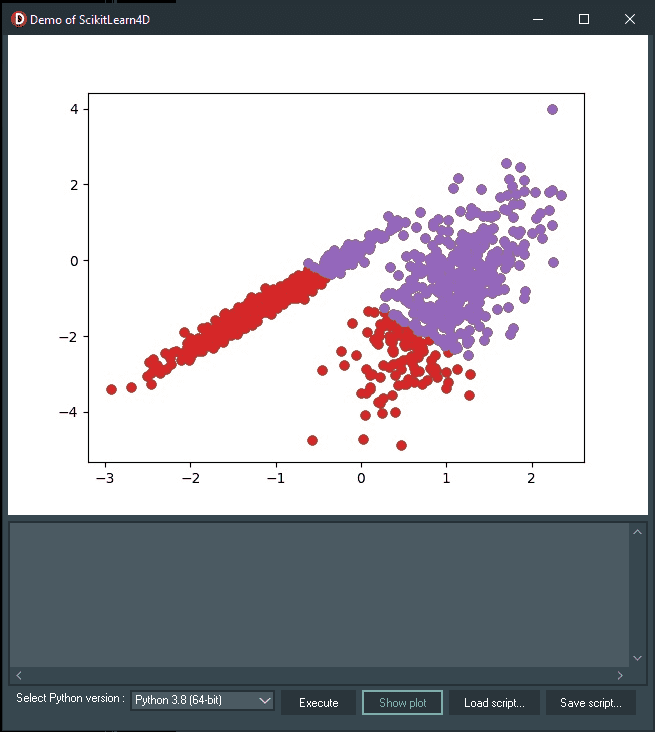
Try out this Agglomerative Clustering example by loading the demo02_agglomerativeClustering.py at runtime by clicking the “Load script…” button, and then “Execute”. Here is the output:
The AgglomerativeClustering class is used to write the demo02_agglomerativeClustering.py, and the main configuration to tune is the “n_clusters” set, an estimate of the number of clusters in the data, e.g. 2.
In this case, we discovered a reasonable grouping result.
What is the BIRCH algorithm?
Quoting Zhang et. al, 1996:
“BIRCH (Balanced Iterative Reducing and Clustering using Hierarchies) incrementally and dynamically clusters incoming multi-dimensional metric data points in an attempt to produce the best quality clustering with the available resources (i. e., available memory and time constraints).
BIRCH can usually find good clustering with a single scan of the data and improve the quality even more with a few additional scans. BIRCH is also the first database clustering algorithm to effectively handle “noise” (data points that are not part of the underlying pattern).
Try out the BIRCH algorithm example by loading demo03_BIRCH.py at runtime by clicking the “Load script…” button, followed by “Execute”.
Running this example, like the previous ones, fits the model on the training dataset and predicts a cluster for each example in the dataset. The points on the scatter plot are then colored by the cluster to which they belong.
Here is the result:

The Birch class is used to write the demo03_BIRCH.py, and the main configuration to tune is the “threshold” and “n_clusters” hyperparameters, the latter of which provides an estimate of the number of clusters.
In this case, using BIRCH, an excellent clustering result is found!
What does DBSCAN mean?
Quoting Ester et. al, 1996:
“DBSCAN, or Density-Based Spatial Clustering of Applications with Noise, is a clustering algorithm that is designed to discover clusters of arbitrary shape by relying on a density-based notion of clusters.
DBSCAN only requires one input parameter and assists the user in determining an appropriate value.“
To try the DBSCAN algorithm example, load demo04_DBSCAN.py at runtime by clicking the “Load script…” button, and then “Execute”. Here is the result:

The DBSCAN class is used to write the demo04_DBSCAN.py, and the main configuration to tune is the “threshold” and “n_clusters” hyperparameters, the latter of which provides an estimate of the number of clusters.
In our case using DBSCAN, a reasonable grouping is found, although more tuning is required.
What is the K-Means machine learning method?
Quoting MacQueen, 1967:
“K-Means is a method for partitioning an N-dimensional population into k sets based on a sample. The process appears to produce partitions that are reasonably efficient in the sense of within-class variance.
…
Furthermore, the k-means procedure is simple to program and computationally economical, making it possible to process very large samples on a digital computer.
…
The problem of “similarity grouping” or “clustering” is perhaps the most obvious application of the k-means process.
The goal of this application is not to find a single, definitive grouping, but rather to help the investigator gain a qualitative and quantitative understanding of large amounts of N-dimensional data by providing him with reasonably good similarity groups.”
To run this k-means clustering example, open the demo05_KMeans.py file by clicking the “Load script…” button and then clicking “Execute.” Here is the result:
The KMeans class is used to write the demo05_KMeans.py, and the main configuration to tune is the “n_clusters” hyperparameter set to the estimated number of clusters in the data.
In this case, a reasonable grouping is found, but the method is less suited to this dataset due to the unequal equal variance in each dimension.
What is the Mini-Batch K-Means machine learning algorithm?
Quoting Béjar, 2013:
“For clustering large datasets, Mini Batch K-means has been proposed as an alternative to the K-means algorithm. The advantage of this algorithm is to reduce the computational cost by not using all the dataset for each iteration but a subsample of a fixed size. This strategy reduces the number of distance computations performed per iteration at the cost of lower cluster quality.”
To try this Mini-Batch K-Means algorithm example, load the demo06_miniBatch_KMeans.py at runtime by clicking the “Load script…” button, and then “Execute”. Here is the output:

The MiniBatchKMeans class is used to write the demo06_miniBatch_KMeans.py, and the main configuration to tune is the “n_clusters” hyperparameter set to the estimated number of clusters in the data.
As usual, running the example fits the model on the training dataset and predicts a cluster for each example in the dataset. The points on the scatter plot are then colored by the cluster to which they belong.
In this case, a result that is almost perfectly equivalent to the standard k-means algorithm is found.
What is the Mean Shift machine learning technique and how is it used?
Quoting Cheng, 1995:
“Mean shift is a non-parametric feature-space mathematical analysis technique for locating density function maxima, also known as a mode-seeking algorithm.
Mean shift is a simple iterative procedure that transforms each data point into the average of data points in its neighborhood. The mean shift algorithm’s generalization makes some k-means-like clustering algorithms into special cases.”
Try out the Mean Shift algorithm by loading demo07_meanShift.py at runtime by clicking the “Load script…” button, followed by “Execute”. Here is the result:

The MeanShift class is used to write the demo07_meanShift.py, and the main configuration to tune is the “bandwidth” hyperparameter.
It resulted in a reasonable set of clusters found in our data.
What is OPTICS cluster analysis?
Quoting Ankerst et. al, 1999:
“OPTICS (Ordering Points To Identify the Clustering Structure) is a cluster analysis algorithm that does not explicitly produce a clustering of a data set, but instead creates an augmented ordering of the database that represents its density-based clustering structure.
This cluster-ordering contains information equivalent to density-based clustering for a wide range of parameter settings. It serves as a versatile foundation for both automatic and interactive cluster analysis.
…
OPTICS clustering is essentially an extended version of the DBSCAN algorithm.”
Experiment with the OPTICS algorithm by loading demo08_OPTICS.py at runtime by clicking the “Load script…” button, and then “Execute”.
Running this example, like the previous ones, fits the model on the training dataset and predicts a cluster for each example in the dataset. The points on the scatter plot are then colored according to the cluster they belong to.
Here is the result:
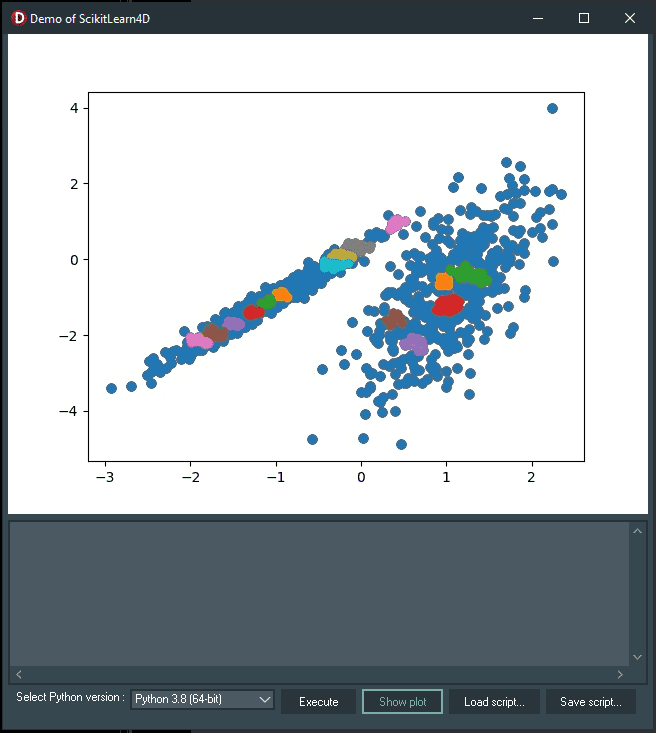
The OPTICS class is used to write the demo08_OPTICS.py, and the main configuration to tune is the “eps” and “min_samples” hyperparameters.
For this example dataset, a reasonable result is not achieved.
What is Spectral Clustering and how is it used in machine learning?
According to Luxburg (2007),
“Compared to “traditional algorithms” like k-means or single linkage, spectral clustering has numerous fundamental advantages.
Spectral clustering results frequently outperform traditional approaches; spectral clustering is very simple to implement and can be solved efficiently using standard linear algebra methods.“
To try the Spectral Clustering algorithm example, load the demo09_spectralClustering.py at runtime by clicking the “Load script…” button and then “Execute”.

The SpectralClustering class is used to write the demo09_spectralClustering.py, and the main configuration to tune is the “n_clusters” hyperparameter used to specify the estimated number of clusters in the data.
Reasonable clusters were discovered in this case.
The Gaussian Mixture machine learning model
According to Reynolds, 2009:
“A Gaussian Mixture Model (GMM) is a parametric probability density function represented as a weighted sum of Gaussian component densities.
GMMs are frequently used in biometric systems as a parametric model of the probability distribution of continuous measurements or features, such as vocal-tract-related spectral features in a speaker recognition system. The iterative Expectation-Maximization (EM) algorithm or Maximum A Posteriori (MAP) estimation from a well-trained prior model is used to estimate GMM parameters from training data.”
To run this Gaussian Mixture Model clustering example, open the demo10_gaussianMixtureModel.py by clicking the “Load script…” button, and then “Execute”. Here is the result:

The GaussianMixture class is used to write demo10_gaussianMixtureModel.py, and the main tuning parameter is the “n_clusters” hyperparameter, which specifies the estimated number of clusters in the data.
In this case, we can see that the clusters were perfectly identified. This is not surprising because the dataset was generated as a mixture of Gaussians.
Such a powerful unsupervised machine learning library, right?
Visit this repository for complete source code.
Are you ready to try these ScikitLearn4Delphi machine learning demos?
Congratulations, now you have learned a lot about scikit-learn: A rich and powerful unsupervised machine learning library, and how you can use Delphi to create a simple yet powerful GUI for it!
We have learned the fundamentals of 10 different clustering algorithms to discover clusters on the data, and from that foundation, you can develop your own AI apps.
For a more Pythonic point of view, you can refer to this article:
If you are looking for other powerful AI libraries, please read this article:
Download a free trial of RAD Studio Delphi today and try out these examples for yourself.
References & Further Readings
Papers
- Thavikulwat, P. (2008). Affinity propagation: A clustering algorithm for computer-assisted business simulations and experiential exercises. In Developments in Business Simulation and Experiential Learning: Proceedings of the Annual ABSEL conference (Vol. 35).
- Ackermann, M. R., Blömer, J., Kuntze, D., & Sohler, C. (2014). Analysis of agglomerative clustering. Algorithmica, 69(1), 184-215.
- Zhang, T., Ramakrishnan, R., & Livny, M. (1996). BIRCH: an efficient data clustering method for very large databases. ACM sigmod record, 25(2), 103-114.
- Ester, M., Kriegel, H. P., Sander, J., & Xu, X. (1996, August). A density-based algorithm for discovering clusters in large spatial databases with noise. In kdd (Vol. 96, No. 34, pp. 226-231).
- MacQueen, J. (1967, June). Some methods for classification and analysis of multivariate observations. In Proceedings of the fifth Berkeley symposium on mathematical statistics and probability (Vol. 1, No. 14, pp. 281-297).
- Béjar Alonso, J. (2013). K-means vs mini batch k-means: A comparison.
- Cheng, Y. (1995). Mean shift, mode seeking, and clustering. IEEE transactions on pattern analysis and machine intelligence, 17(8), 790-799.
- Ankerst, M., Breunig, M. M., Kriegel, H. P., & Sander, J. (1999). OPTICS: Ordering points to identify the clustering structure. ACM Sigmod record, 28(2), 49-60.
- Von Luxburg, U. (2007). A tutorial on spectral clustering. Statistics and computing, 17(4), 395-416.
- Reynolds, D. A. (2009). Gaussian mixture models. Encyclopedia of biometrics, 741(659-663).
Links
- https://towardsdatascience.com/unsupervised-machine-learning-affinity-propagation-algorithm-explained-d1fef85f22c8/
- https://docs.rapidminer.com/latest/studio/operators/modeling/segmentation/agglomerative_clustering.html/
- https://machinelearningmastery.com/clustering-algorithms-with-python/
- https://en.wikipedia.org/wiki/DBSCAN/
- https://www.analyticsvidhya.com/blog/2019/08/comprehensive-guide-k-means-clustering/
- https://thecleverprogrammer.com/2021/09/10/mini-batch-k-means-clustering-in-machine-learning/
- https://analyticsindiamag.com/hands-on-tutorial-on-mean-shift-clustering-algorithm/
- https://towardsdatascience.com/spectral-clustering-aba2640c0d5b/
- https://towardsdatascience.com/gaussian-mixture-models-d13a5e915c8e/




